 Members of the Blazer Data Wranglers team, who analyzed COVID-19 disease burden and comorbidities in African American patients at UAB during the two-day hackathon.Can data science help solve COVID-19 challenges? And what can you accomplish with 48 hours, a massive amount of data and freedom to explore fresh ideas? Judging by the entries in UAB’s COVID–19 Data Science Hackathon, the answers seem to be “yes” and “quite a bit.”
Members of the Blazer Data Wranglers team, who analyzed COVID-19 disease burden and comorbidities in African American patients at UAB during the two-day hackathon.Can data science help solve COVID-19 challenges? And what can you accomplish with 48 hours, a massive amount of data and freedom to explore fresh ideas? Judging by the entries in UAB’s COVID–19 Data Science Hackathon, the answers seem to be “yes” and “quite a bit.”
On June 15 and 16, following an all-day bootcamp to get them up to speed on the tools and data available, 38 participants on 10 teams tore into seriously ambitious projects. Nearly half of the group were students; the rest were faculty and staff from UAB and several other institutions nationwide. By the end of the week, the teams demonstrated their results in presentations over Zoom.
A hackathon — a hacking marathon — is “an intensive collaboration on software development to advance knowledge and innovation,” said Amy Wang, M.D., associate professor of medicine and scientist in the UAB Informatics Institute, which sponsored the event along with the Center for Clinical and Translational Science and the AI.Med Lab at UAB. In 2018, the Informatics Institute convened the first UAB data-science hackathon with only four teams. “Getting everyone in the same room, giving them food and helping them innovate with the right biocomputing support” helped launch the first version of the Informatics Institute’s U-BRITE team science platform, Wang said.
| Watch all the COVID-19 Data Science Hackathon presentations. |
This spring, with COVID-19 affecting research productivity across UAB, U-BRITE emerged as a digital platform in which investigators can regroup from their homes to solve important problems, said Jake Chen, Ph.D., professor of genetics, computer science and biomedical engineering at UAB and chief bioinformatics officer of the UAB Informatics Institute. “The COVID-19 Data-driven Medicine Hackathon is a natural convergence of several parallel efforts,” Chen said. “Many trainees and professionals are curious about biomedical data science. Our U-BRITE development added new features and capabilities to serve domain-specific biomedical research as of version 2.0, which had just launched. And COVID-19 forced us to bring people, technology and science together on a web portal as quickly as we can, culminating in the hackathon showcase competition.”
Maps for COVID travel, simulations and machine learning insights
This time, no one was in the same room — participants did all their collaboration through Zoom and other digital tools in a “virtual hackathon.” But this didn’t appear to hamper anyone’s creativity or passion to work together across a range of disciplines that included computer science, biomedical informatics, engineering, clinical practice, biomedical sciences and health services.
One team developed a route-finder, along the lines of Google Maps, that enables travelers to plan refueling and overnight stops so as to avoid counties with sizable COVID-19 outbreaks. Multiple groups used de-identified data from COVID-19 hospitalizations at UAB to gain insight on patients at severe risk of the disease. Others trained machine-learning models on national datasets to identify vulnerable populations and validate targets for drugs and adapted the evolutionary simulation Game of Life to model the spread of COVID-19. (See a list of all the projects athttp://ubrite.informatics.uab.edu/covid19/.)
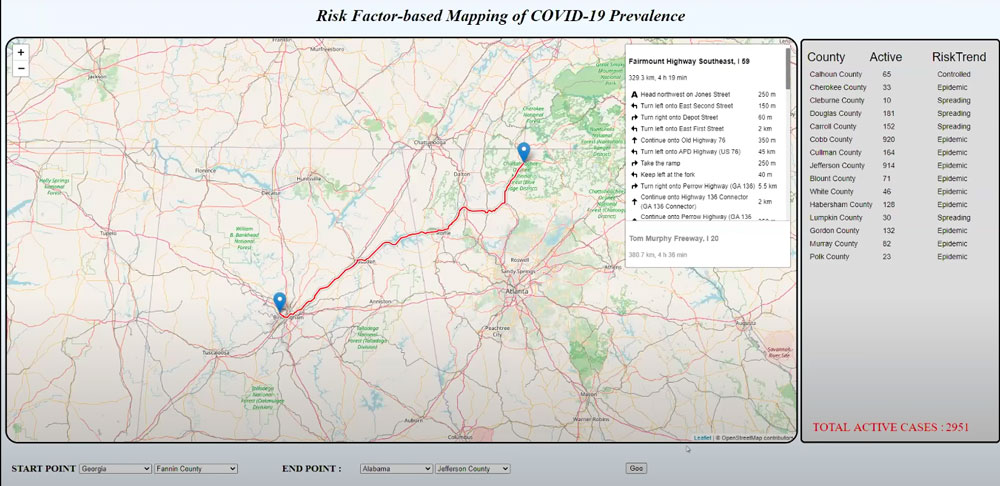 Team 2's "Risk factor-based mapping of COVID-19 prevalence" project produced this interactive map that takes county-level disease data into account when calculating driving directions. Watch the full presentation here.
Team 2's "Risk factor-based mapping of COVID-19 prevalence" project produced this interactive map that takes county-level disease data into account when calculating driving directions. Watch the full presentation here.
“I am pleased that the teams appear to have found the U-BRITE platform to be a productive place to hack together and seemed to have had a good time while doing it,” said Jelai Wang, IT architect at the Informatics Institute, who led the pre-hackathon bootcamp. “Hackathon planning and IT infrastructure development can be a lot of hard work and stress behind the scenes, so for us to set out to support and empower these hackers with COVID-19 data and tools within U-BRITE and then actually see teams organize around interesting questions, hack under time pressure and produce results is quite rewarding.”
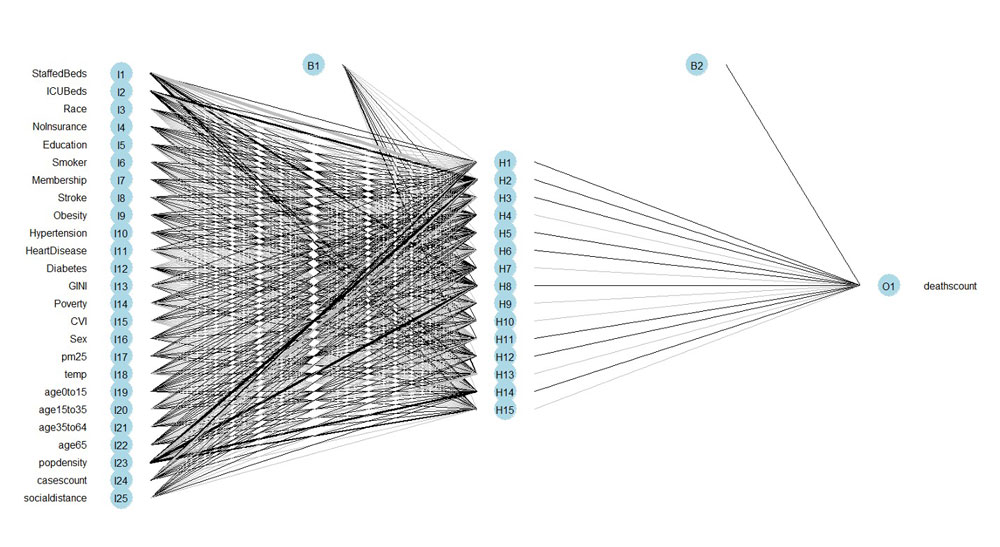 Slide from Team 8's project "Predicting fatalities and spatial vulnerabilities due to COVID-19 in the United States." Watch the full presentation here.
Slide from Team 8's project "Predicting fatalities and spatial vulnerabilities due to COVID-19 in the United States." Watch the full presentation here.
The hackathon had two main goals, Chen explained. One was to develop solutions using datasets stored or managed in U-BRITE. The other was to “promote learning and collaboration about new tools, software and technologies for data science,” he said. COVID-19 represents “a unique opportunity for us in the School of Medicine,” he noted. “The science is so new and so many teams worldwide are generating a tidal wave of data — epidemiology, clinical, social, imaging, genomic data for example — that a lot of the traditional experts don’t even know how to fight it. This is a rare opportunity for those who are skilled with big data and AI tools to join the team and lead.”
Winning projects
Three winning projects were selected by a panel of nine judges.
First prize went to Curtis Hendrickson, a research associate with the Center for Clinical and Translational Science, whose Novel2Global project developed an automated pipeline to compare patient-specific viral genomes from patients at UAB Hospital with global reference strains. (Watch Hendrickson’s Novel2Global presentation.) This could allow for “genomic epidemiology” to find virus clusters in the community and transmission within the hospital. “We thought we should have these tools up and running here at UAB,” said Hendrickson, who worked with Elliot Lefkowitz, Ph.D., a professor in the Department of Microbiology and director of Informatics at the CCTS with expertise in viral phylogeny. Hendrickson noted that similar work from researchers at Yale University was published in the journal Cell in May. “It’s quite impressive that something that we’re taking on at a hackathon was in Cell only a month ago,” Hendrickson said.
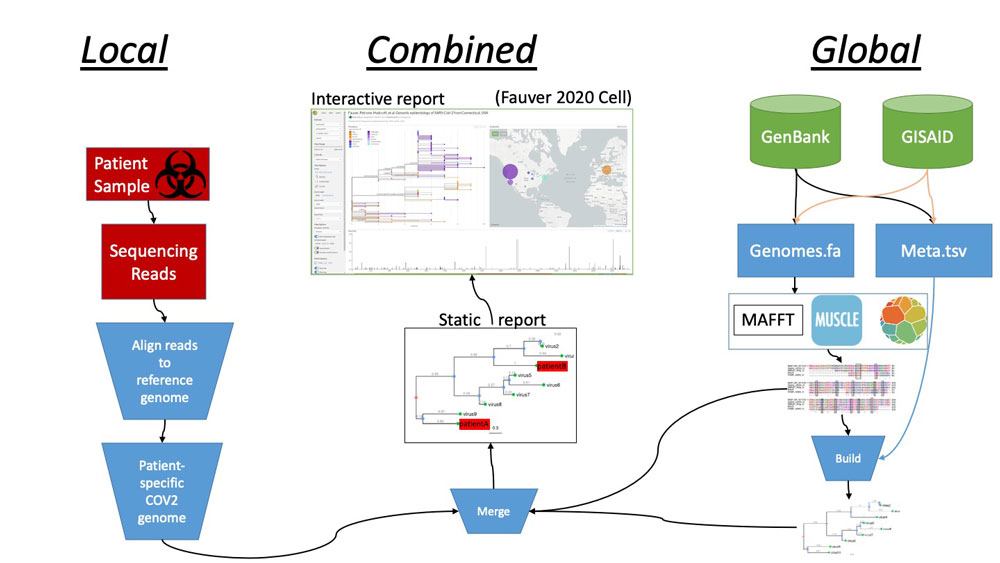 A slide from Curtis Hendrickson's winning Novel2Global project, which aimed to automate comparison of viral genomes seen in patients at UAB with global reference strains to enable genomic epidemiology. Watch the full presentation here.
A slide from Curtis Hendrickson's winning Novel2Global project, which aimed to automate comparison of viral genomes seen in patients at UAB with global reference strains to enable genomic epidemiology. Watch the full presentation here.
Second prize was awarded to the COVID-19 SIMULATE team, which developed a network-based epidemiologic model to simulate transmission of COVID-19 across several levels, including families, counties and states. (Watch the COVID-19 SIMULATE team’s presentation.) Team members were Zongliang Yue, a doctoral candidate in genetics, genomics and bioinformatics; Eric Zhang, a doctoral candidate in pathobiology and molecular medicine; Josep Rubio Pique, a student in the M.S. in Data Science program in the Department of Computer Science; and mentors Da Yan, Ph.D., assistant professor in computer science; and Jake Chen, Ph.D., professor in the departments of Genetics, Computer Science and Biomedical Engineering.
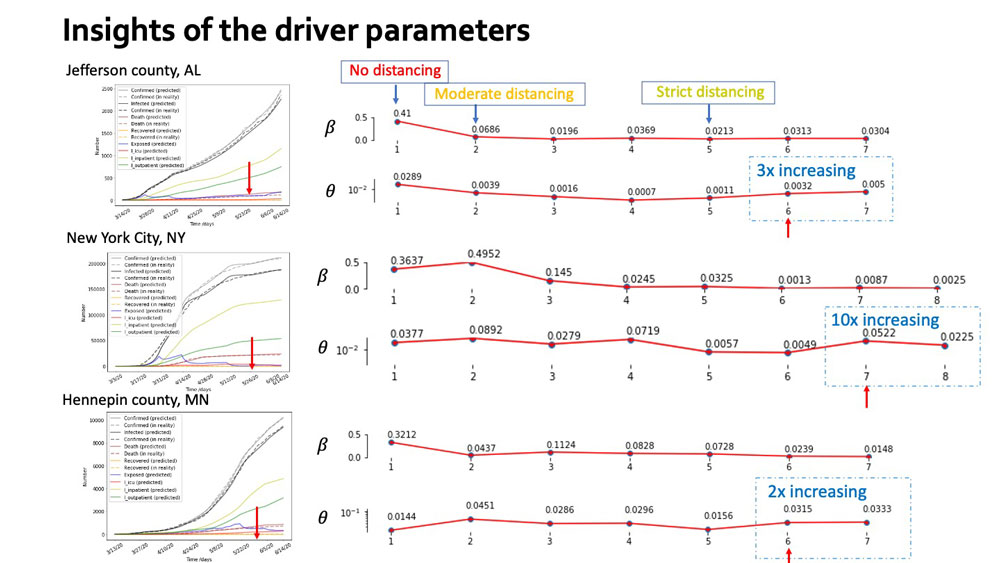 Slide from the COVID-19 SIMULATE team's presentation on its network-based epidemiologic model of disease transmission across multiple levels. Watch the full presentation here.
Slide from the COVID-19 SIMULATE team's presentation on its network-based epidemiologic model of disease transmission across multiple levels. Watch the full presentation here.
Third prize went to the RICO (RIsk of COvid) team, which adapted credit scorecard models used in the financial industry to create a functioning web app that advises users whether or not they should be tested for COVID-19 based on their symptoms. (Watch the RICO team’s presentation.) Team members were: Tarun Mamidi, doctoral candidate in genetics, genomics and bioinformatics; Thi Tran-Nguyen, Ph.D., committee chair and data scientist for UAB’s U-BRITE/COVID-19 Knowledge Curation Taskforce and a graduate of the doctoral program in immunology; Ryan Melvin, Ph.D., assistant professor of anesthesiology and perioperative medicine; and mentor Elizabeth Worthey, Ph.D., associate professor of pediatric hematology and oncology.
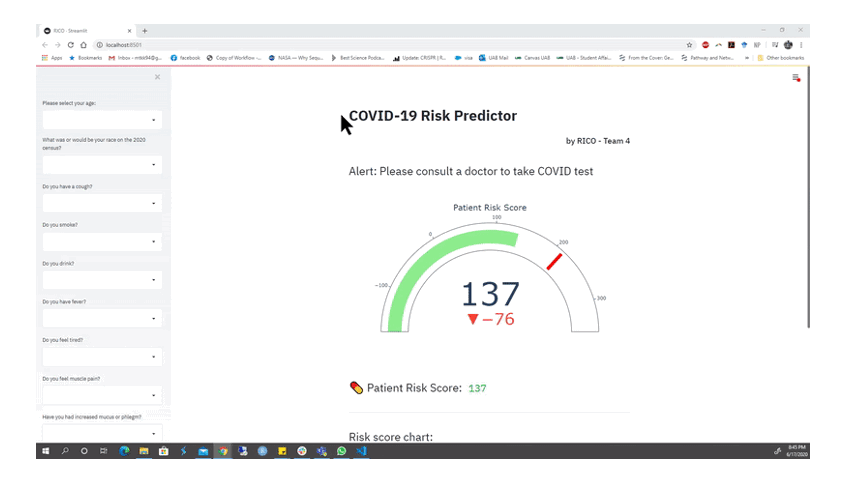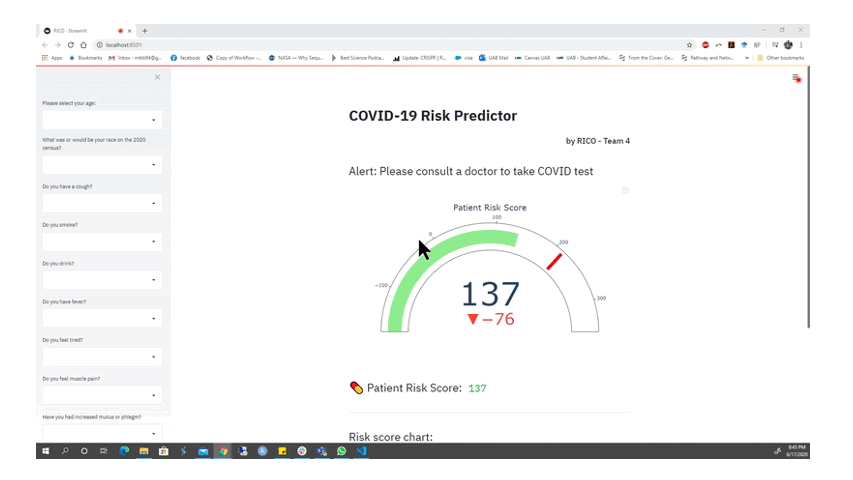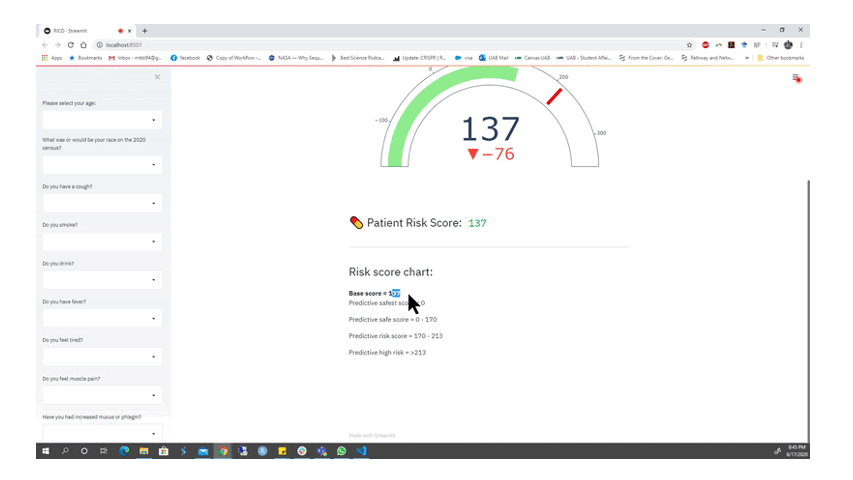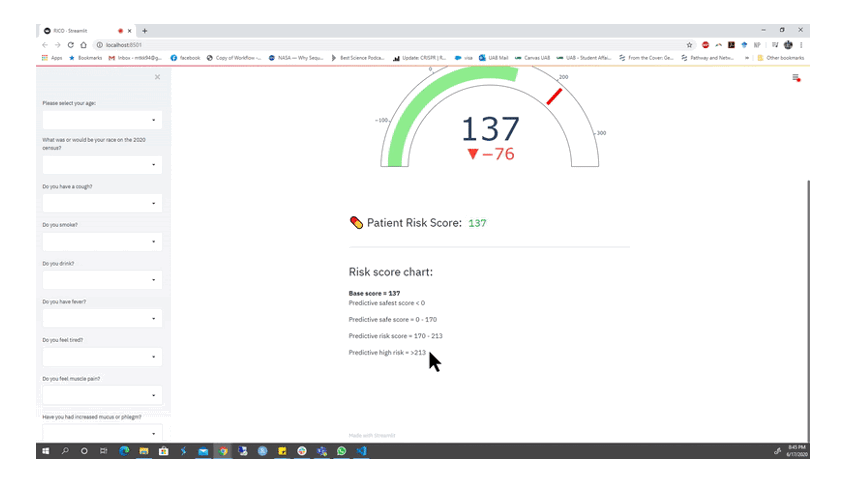 Animation from the RICO team depicting part of their functioning web app for alerting members of the public if they should seek a COVID-19 test. Watch the full presentation here.
Animation from the RICO team depicting part of their functioning web app for alerting members of the public if they should seek a COVID-19 test. Watch the full presentation here.

“This was my second hackathon. The first time I just watched; this second time I was able to throw out my idea and to my amazement others rallied around it…. I learned Cheaha, GitLab and R Studio in the weeks leading up to the hackathon, and our team continues to work together to write about what we studied.”
— Hope Gray, graduate research assistant and doctoral student in health services administration at UABProjects moving forward
The hackathon is just the beginning for many of the projects presented.
Hendrickson’s idea for Novel2Global came from a project that he, Lefkowitz and Liam Van Der Pol in the CCTS Informatics Group are working on with Sixto Leal, M.D., assistant professor in the Department of Pathology. That project, funded by the School of Medicine’s urgent COVID-19 research grants, will sequence the transcriptomes of patients with COVID-19 and the genomes of the SARS-CoV-2 viruses infecting them at the same time. That will allow Leal to “compare genes active in the patients with different outcomes, and the genomes of the virus, allowing us to look at both the host response and the virus genetics in order to try and understand why some patients have a more severe response, and thus understand the disease process better,” Hendrickson said. But the hackathon “also provided an opportunity to do more,” he noted. “The global phylogenic placement of the viral genomes is not an immediate priority for Dr. Leal’s project, but will add to the planned analysis and make it richer and more informative. It will also, then, be available for anyone else doing a similar project, as all the code will be publicly available, and we’ll be available to help people use it on their own data. (Interested parties should contact Lefkowitz, Hendrickson said.)
Meanwhile, the RICO team is expanding the data sources used in their original model and implementing new, more powerful machine-learning algorithms. They also are exploring the potential of linking their model directly to symptom trackers already in use.
The winners were invited to also present their work at the 19th International Workshop on Data Mining in Bioinformatics in August 2020, in which Da Yan, Ph.D., an assistant professor in UAB’s computer science department, and Chen are organizers. And all participants were invited to submit their work for consideration in a special issue of the journal Frontiers on Artificial Intelligence being organized by Chen.
The hackathon itself will help improve U-BRITE, Jelai Wang said. “We received valuable feedback from hackathon participants and learned more about what parts of U-BRITE worked well and what could be improved,” he said. “This will feed into our plans for future development.”
‘A lot of joy at the end’
Several participants noted that they valued the focus brought by the hackathon format. “This was a wonderful experience,” said Hope Gray, a graduate research assistant and doctoral student in health services administration at UAB, whose group studied COVID-19 disease burden and comorbidities in African American patients using data from UAB Hospital. Gray said this was her second hackathon. “The first time I just watched; this second time I was able to throw out my idea and to my amazement others rallied around it.” Her group included students from Tuskegee University and UAB and faculty from Johns Hopkins University, the University of Maryland and UAB. “I learned Cheaha, GitLab and R Studio in the weeks leading up to the hackathon, and our team continues to work together to write about what we studied.”
“It was an opportunity for us to flesh out some ideas that we already had before we knew about the hackathon,” said Jason White, a research core lab manager and student at Tuskegee. His group, which included other Tuskegee students, a UAB student and a faculty member at the University of Wisconsin, examined racial disparities in COVID-19 using gene-expression data. “It can be hard to find the data you need to answer the questions you have but this gave us the opportunity to drop other things and focus,” White said. “And we were successful in finding data we could use going forward.”
Gargya Malla, a doctoral student in epidemiology at UAB, described herself as a clinician, not a technical person. “I really liked the idea that I could work with people who are not in public health but still move forward a public health idea,” Malla said. Her team worked on understanding the impact of social determinants of health on COVID-19 outcomes, with one member doing Python coding, another focused on machine-learning and another on data visualizations. “It was a very stressful 48 hours which gave a lot of joy by the end,” Malla said. “Thanks to the organizers for allowing us to collaborate with people who have nothing in common.”
That is, except for a shared desire to make an impact on COVID-19. “None of us, except for Bob [Kimberly, M.D., CCTS director] and a few clinicians are fighting on the front lines,” Chen said. “Are we even relevant? I think we all can become relevant by becoming codebreakers.” Computer scientists such as Alan Turing transformed the war effort by cracking enemy codes, Chen said. “That is very much analogous to what we are doing in the hackathon.”
Hendrickson said there was “no secret sauce” to his success. “It's having everybody putting in the energy to make this event happen that gives us the opportunity to do something interesting and special, and gives us all these resources that we have access to — the mentors and the infrastructure — and then giving us a chunk of time to work together,” Hendrickson said. “That makes a huge difference to moving projects forward. It's not our special sauce. It's the fact that everybody at the institution worked so hard to create this space for us to flourish in.”
Not the only opportunity
This isn’t the only opportunity to join the fight, Chen noted. The Informatics Institute and the CCTS are leading UAB’s participation in the NIH-sponsored National COVID Cohort Collaborative (N3C), which aims to build a centralized national data resource for the research community can use to study COVID-19 and identify potential treatments, according to the N3C website. “If you are interested in taking part, contact me, [Precision Medicine Institute director] Matt Might or [Informatics Institute director] Jim Cimino,” Chen said.
