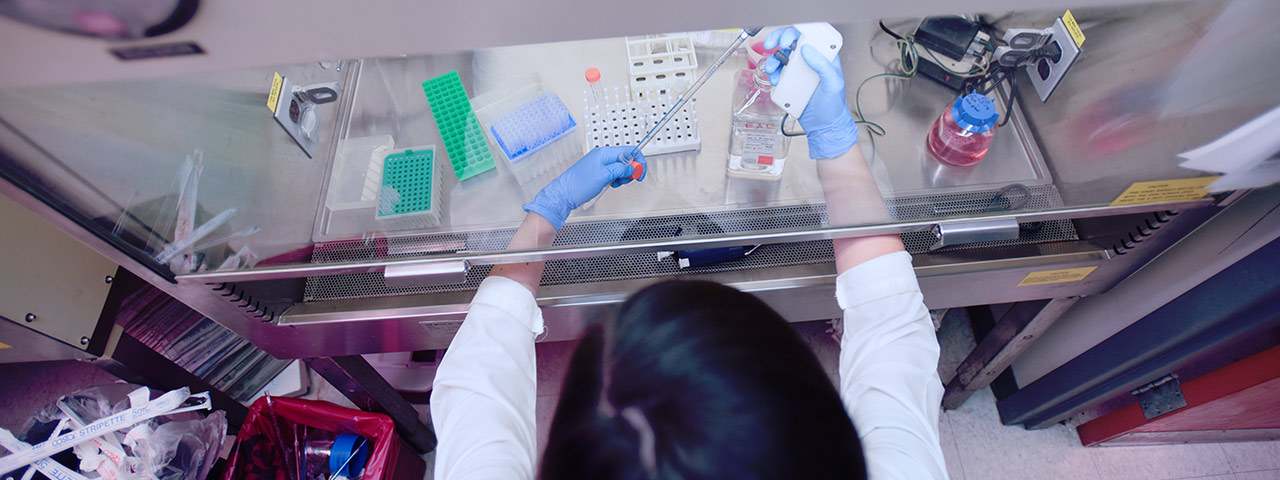
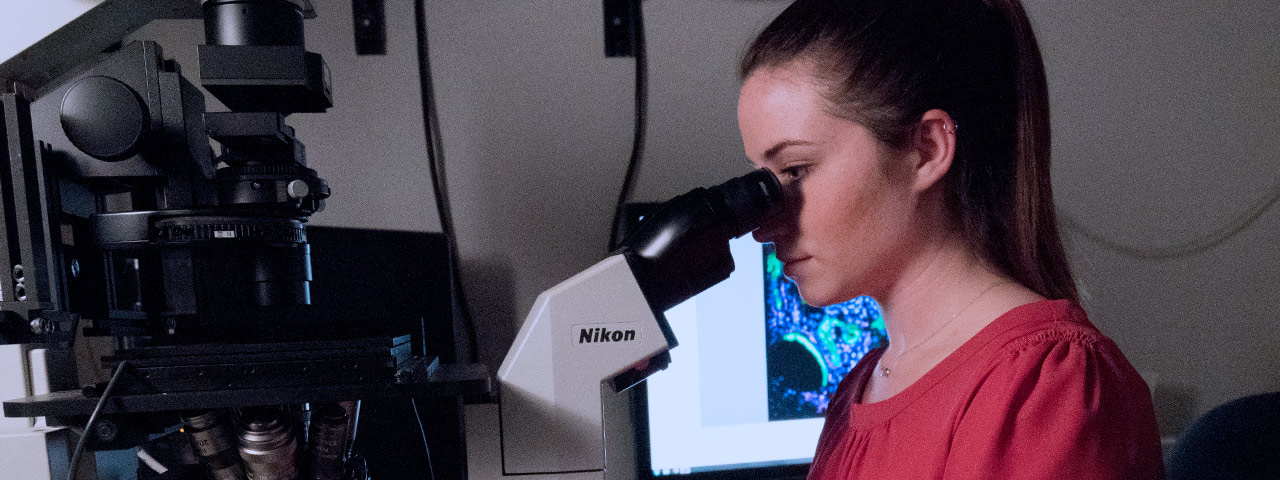
Graduate Biomedical Sciences at the University of Alabama at Birmingham offers eight interdisciplinary Ph.D. training themes. We are deeply committed to seeking, exploring, innovating, and ultimately discovering long-sought causes of disease, life-saving therapies, and world-changing breakthroughs. Join us as we change the future through discovery.
Graduate Biomedical Sciences at UAB — The University of Alabama at Birmingham
Innovation in Modern Biomedicine